A benchmark of batch-effect correction methods for single-cell RNA sequencing data | Genome Biology | Full Text

Evaluation of batch effect correction methods for... - Lauren Sanders - BOSC - Abstract - ISMB 2022 - YouTube
Performance assessment of seven batch-effect correction methods on cell... | Download Scientific Diagram

Scaling Method for Batch Effect Correction of Gene Expression Data Based on Spectral Clustering | Bentham Science

NormAE: Deep Adversarial Learning Model to Remove Batch Effects in Liquid Chromatography Mass Spectrometry-Based Metabolomics Data | Analytical Chemistry

Correcting batch effects in single-cell RNA sequencing data by matching mutual nearest neighbours | bioRxiv

BatchServer: A Web Server for Batch Effect Evaluation, Visualization, and Correction | Journal of Proteome Research
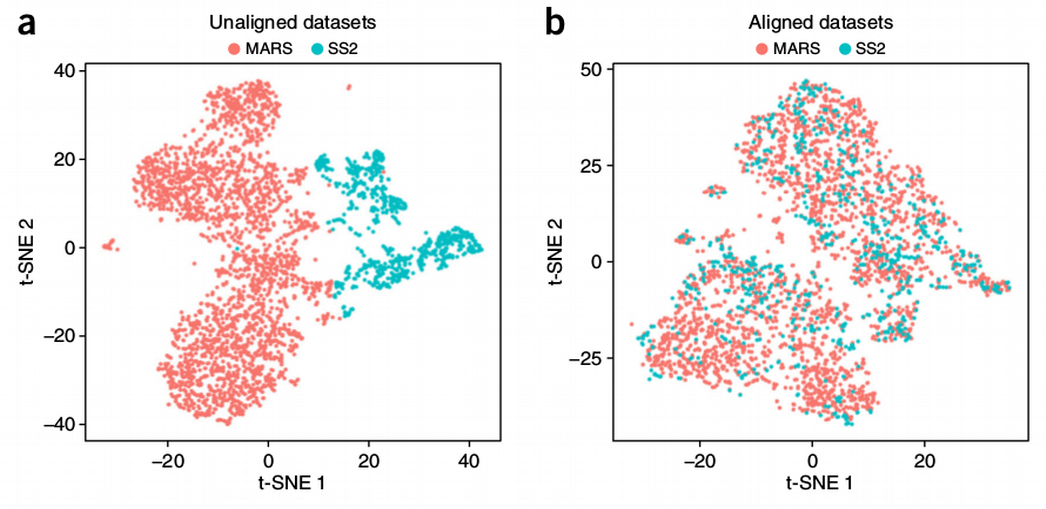
How to Batch Correct Single Cell. Comparing batch correction methods for… | by Nikolay Oskolkov | Towards Data Science

A comparison of methods accounting for batch effects in differential expression analysis of UMI count based single cell RNA sequencing - ScienceDirect

Comparisons of data integration methods for batch effect correction for... | Download Scientific Diagram
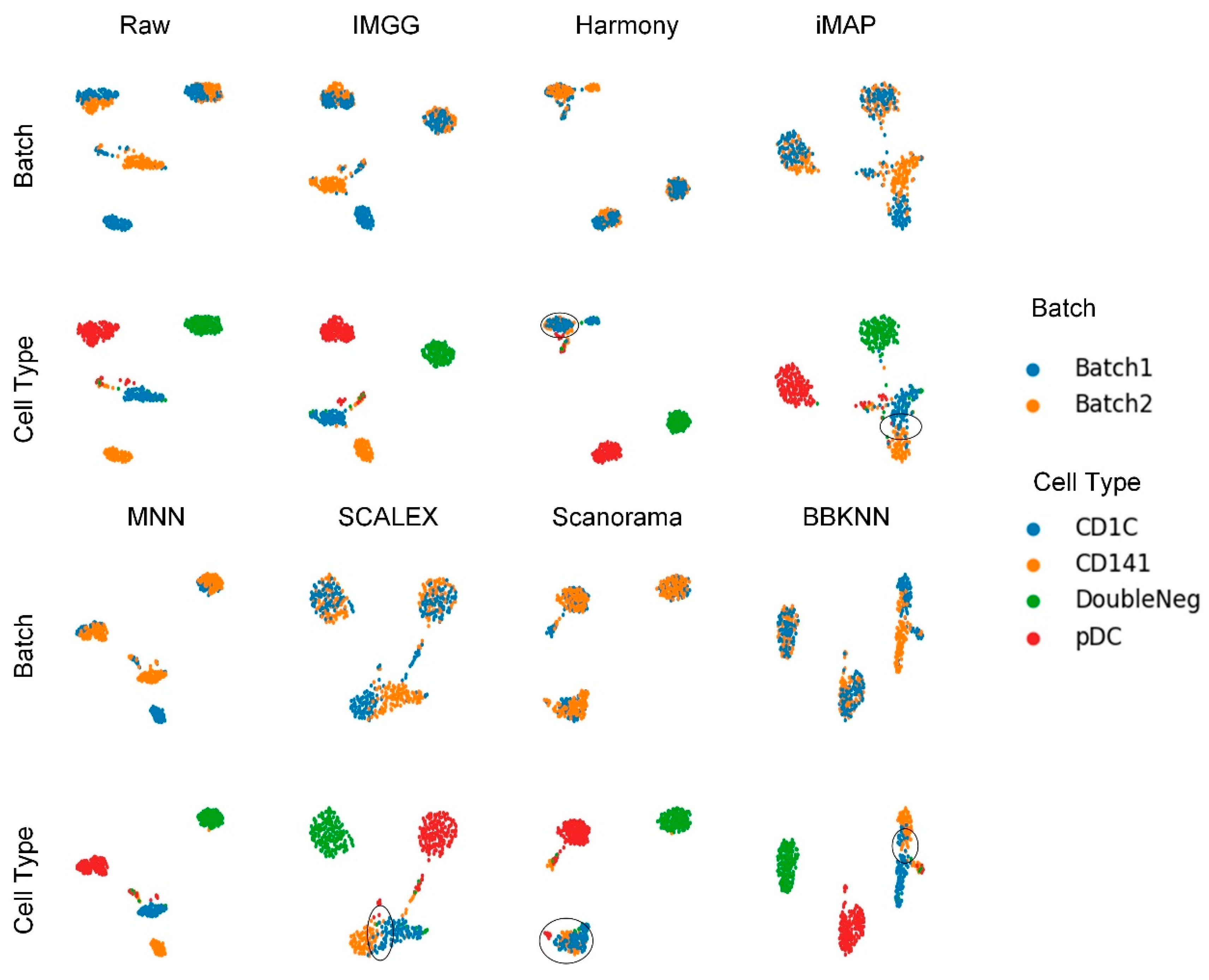

IJMS | Free Full-Text | IMGG: Integrating Multiple Single-Cell Datasets through Connected Graphs and Generative Adversarial Networks

Batch effect reduction of microarray data with dependent samples using an empirical Bayes approach (BRIDGE)

A benchmark of batch-effect correction methods for single-cell RNA sequencing data | Genome Biology | Full Text

Qualitative evaluation of 14 batch-effect correction methods using UMAP... | Download Scientific Diagram
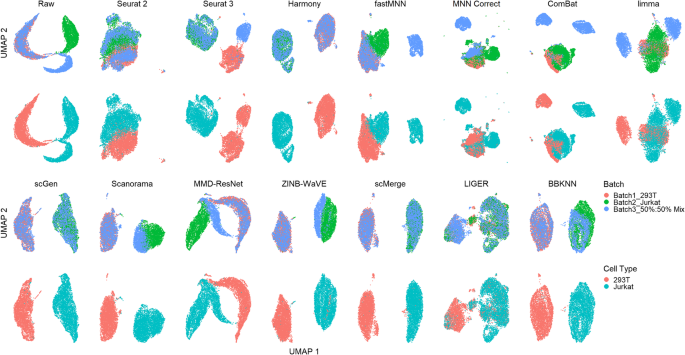
A benchmark of batch-effect correction methods for single-cell RNA sequencing data | Genome Biology | Full Text

Evaluation of batch effect correction methods for... - Lauren Sanders - BOSC - Poster - ISMB 2022 - YouTube

A benchmark of batch-effect correction methods for single-cell RNA sequencing data | Genome Biology | Full Text
GitHub - JinmiaoChenLab/Batch-effect-removal-benchmarking: A benchmark of batch-effect correction methods for single-cell RNA sequencing data
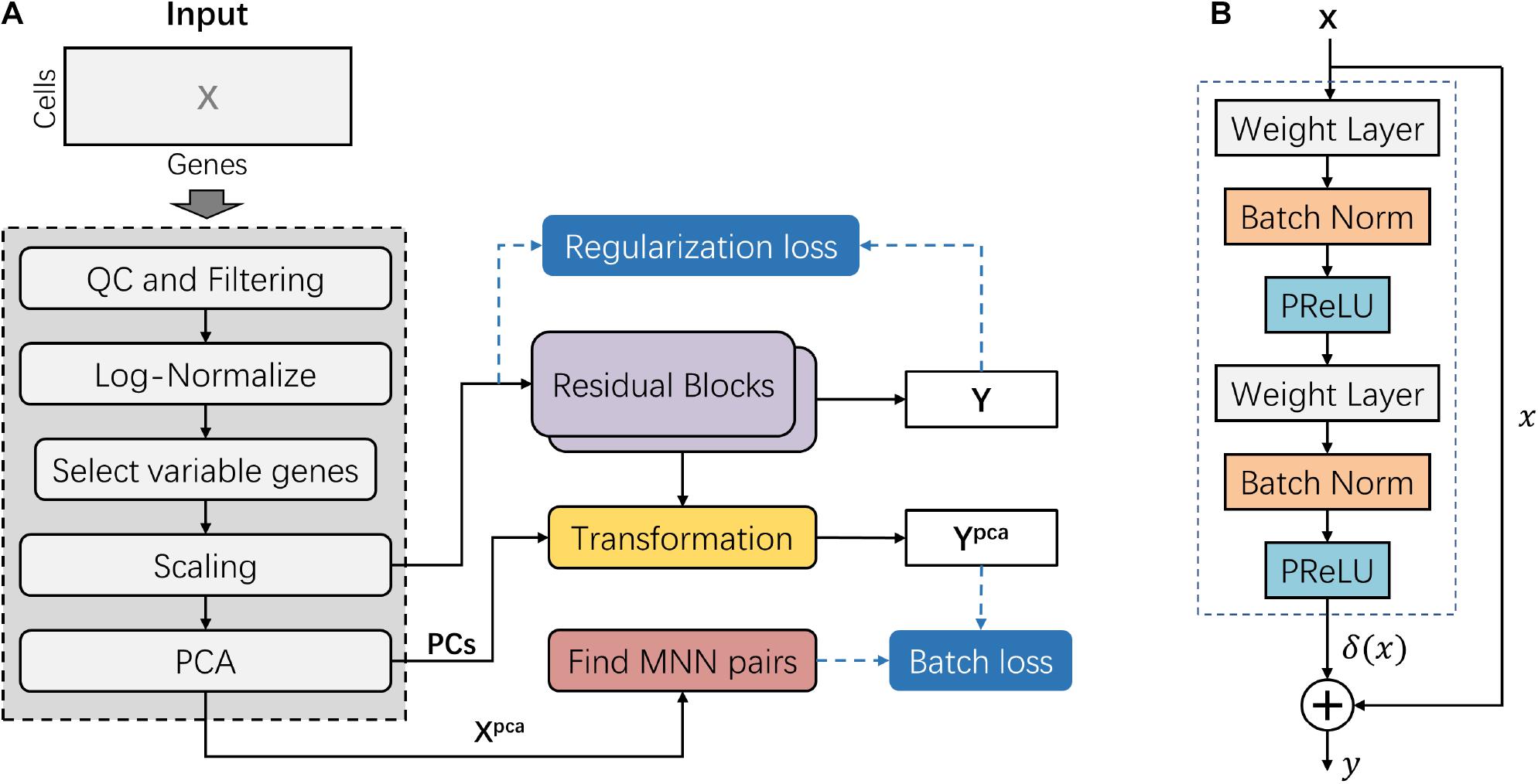
Frontiers | deepMNN: Deep Learning-Based Single-Cell RNA Sequencing Data Batch Correction Using Mutual Nearest Neighbors